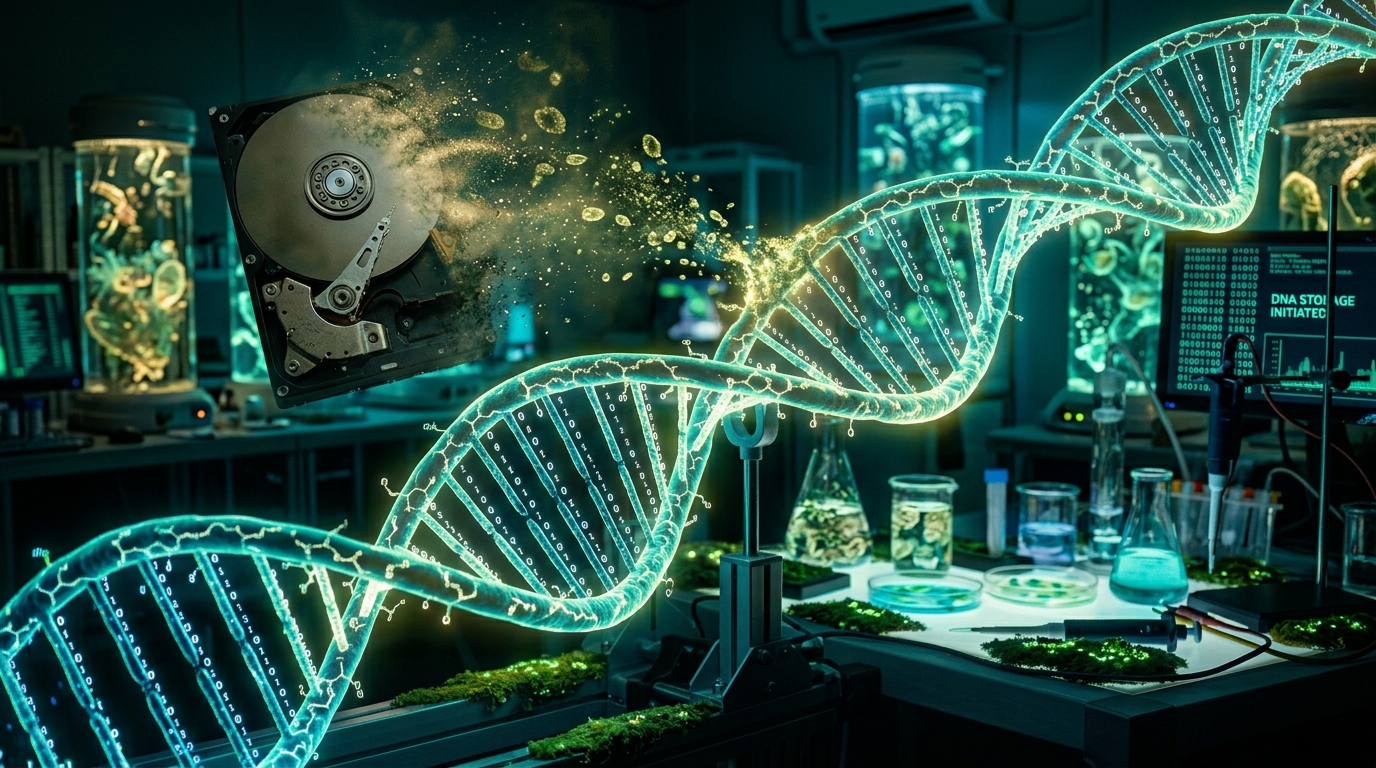
DNA Data Storage: How Biology Could Solve the World’s Exploding Data Crisis
DNA data storage — encoding digital information in synthetic DNA molecules — has advanced from academic curiosity to a technology attracting hundreds of millions in investment and producing working prototypes that could eventually solve the world’s exploding data storage crisis. The global datasphere is projected to reach 175 zettabytes by 2025 and 660 zettabytes by 2030. Current storage technologies — hard drives, solid-state drives, and magnetic tape — face physical limits on density, energy consumption, and longevity. DNA offers a theoretical storage density 1 million times greater than today’s hard drives, durability measured in thousands of years rather than decades, and zero energy consumption for data at rest. The technology is real, the encoding science works, and the remaining challenge is cost and speed.
How DNA Storage Works
DNA stores information in sequences of four nucleotide bases: adenine (A), thymine (T), guanine (G), and cytosine (C). Digital data, which is inherently binary (sequences of 0s and 1s), is mapped onto these four bases using encoding schemes — the simplest being A=00, T=01, G=10, C=11, though practical encoding uses more sophisticated mappings that include error-correcting codes. The encoded nucleotide sequence is then synthesized into physical DNA molecules using standard molecular biology techniques.
To write data, a digital file (an image, a document, a database) is converted to a binary sequence, mapped to nucleotide bases through the encoding algorithm, and then physically synthesized as DNA strands. Multiple copies of each strand are produced for redundancy. The DNA is typically stored in dehydrated form — either as a dried pellet or encapsulated in silica nanoparticles for long-term preservation.
To read data, the stored DNA is sequenced (its nucleotide sequence is determined using DNA sequencing technology), the nucleotide sequence is decoded back to binary using the reverse of the encoding algorithm, and the original digital file is reconstructed. Error-correcting codes embedded in the encoding ensure that sequencing errors (which are inevitable) can be detected and corrected, maintaining data integrity.
The theoretical information density of DNA is staggering. One gram of DNA can store approximately 215 petabytes (215 million gigabytes) of data. For perspective, that’s enough to store all data currently generated by the human race in approximately one day. The entire world’s data could theoretically be stored in a few kilograms of DNA. This density is six orders of magnitude (1 million times) greater than the densest conventional storage technology.
Why DNA Storage Matters: The Data Deluge
The motivation for DNA storage isn’t futuristic novelty — it’s a practical response to a concrete infrastructure problem. The world generates approximately 330 exabytes of data per year, and much of it must be retained for regulatory, scientific, cultural, or business reasons. Archival data — data that must be kept but is rarely accessed — represents 60-80% of stored data in most organizations. This archival data currently sits on magnetic tape (the cheapest conventional archival medium at approximately $5-$10 per terabyte) in climate-controlled tape libraries that consume significant physical space and continuous electricity.
Magnetic tape has a rated lifespan of 15-30 years, after which data must be migrated to new tape (a process called “data migration” that is expensive, error-prone, and must be repeated every few decades indefinitely). Hard drives and SSDs have shorter active lifespans. DNA, by contrast, can survive for thousands of years when stored properly — DNA extracted from 700,000-year-old horse fossils has been successfully sequenced, demonstrating remarkable longevity under far less controlled conditions than a storage facility.
The combination of extreme density (eliminating physical storage space), zero energy for data at rest (no climate control needed if stored in silica capsules), and extraordinary longevity (no periodic migration) makes DNA potentially the most economical medium for archival storage over long timescales. The economic threshold where DNA storage becomes cheaper than tape (accounting for total cost of ownership including space, energy, and migration) is estimated to be within 5-10 years, depending on the trajectory of DNA synthesis cost reductions.
Technical Milestones and Demonstrations
Researchers and companies have demonstrated increasingly sophisticated DNA storage capabilities. Microsoft Research and the University of Washington have been collaborating on DNA storage since 2015 and have demonstrated the storage and perfect retrieval of over 200 MB of data — including the Universal Declaration of Human Rights, a music video, Shakespeare’s Sonnets, and a database — in DNA. Their system uses a sophisticated indexing scheme that enables random access (reading specific data without sequencing the entire archive), addressing one of DNA storage’s key practical requirements.
In 2024, Catalog DNA (now acquired by Seagate) demonstrated a DNA storage system that could write data at a rate of 4 megabits per second — orders of magnitude faster than previous demonstrations, though still vastly slower than conventional storage. Their approach uses a library of pre-synthesized DNA molecules that are combinatorially assembled to encode data, rather than synthesizing each molecule from scratch (the slower, conventional approach). This combinatorial approach trades some storage density for dramatically improved write speed.
Twist Bioscience, a leading DNA synthesis company, has partnered with Microsoft on a large-scale DNA storage development program and announced the production of the first commercially available DNA storage media. Their silicon-based DNA synthesis platform can produce millions of unique DNA sequences in parallel, and continued scaling of this technology is expected to further reduce the cost per base of DNA synthesis.
IARPA (the US Intelligence Advanced Research Projects Activity, the intelligence community’s research arm) has funded DNA storage research through its Molecular Information Storage (MIST) program, reflecting the intelligence community’s interest in ultra-dense, long-term archival storage for classified materials. The MIST program set ambitious targets: writing 1 terabyte of data to DNA within 24 hours at a cost below $1,000 — targets that are not yet achieved but are driving research toward practical scalability.
The Cost Barrier and Path Forward
The primary barrier to practical DNA storage is cost, driven overwhelmingly by the cost of DNA synthesis (writing). Current commercial DNA synthesis costs approximately $0.05-$0.10 per nucleotide base. Since each nucleotide encodes 2 bits of data, storing one megabyte requires approximately 4 million bases at a cost of approximately $200,000-$400,000 per megabyte. For comparison, storing one megabyte on magnetic tape costs approximately $0.00001 (one thousandth of a cent). The cost gap is approximately 10 billion to 1.
However, DNA synthesis costs have been decreasing exponentially over the past two decades — a trajectory sometimes compared to Moore’s Law for semiconductors. The cost per base has dropped approximately 1,000-fold since 2000, and continued improvements in synthesis technology (semiconductor-based synthesis, enzymatic synthesis replacing chemical synthesis, massively parallel synthesis) are projected to continue this trend. Researchers estimate that DNA storage becomes economically competitive with tape at a synthesis cost of approximately $0.001 per base — roughly a 50-100x reduction from current commercial pricing.
Reading (sequencing) DNA is already relatively cheap and fast, thanks to the genomics industry’s massive investment in sequencing technology over the past 20 years. Nanopore sequencing technology (from Oxford Nanopore Technologies) can sequence DNA in real-time using portable devices, providing read capabilities that are already practical for data retrieval. Illumina’s sequencing platforms provide higher accuracy at lower per-base cost for batch reading. The read side of DNA storage is approaching practical viability; it’s the write side (synthesis) that requires the cost breakthroughs.
Enzymatic DNA synthesis — using DNA polymerase enzymes to add nucleotide bases one at a time under controlled conditions — is a promising alternative to traditional chemical synthesis that could dramatically reduce costs. Companies including DNA Script, Ansa Biotechnologies, and Nuclera are developing enzymatic synthesis platforms that operate in aqueous (water-based) conditions at room temperature, eliminating the harsh chemicals and waste products of traditional synthesis. Enzymatic synthesis also has the potential for very high parallelism, which would increase throughput by orders of magnitude.
Applications and Early Adopters
The initial commercial applications for DNA storage will target the archival tier — data that must be retained for decades to centuries but is rarely accessed. Government archives (national records, census data, intelligence archives), cultural preservation (digitized museum collections, film archives, historical records), scientific data (genomic databases, astronomical observations, climate records), and financial records (regulatory retention requirements of 7-30+ years) are the use cases where DNA’s density, longevity, and zero-maintenance properties provide the strongest value proposition.
The film industry has shown particular interest. The Academy of Motion Picture Arts and Sciences estimates that storing a single master copy of a Hollywood feature film in digital-ready format requires approximately 2 petabytes — and the industry produces hundreds of films per year, all of which must be archived indefinitely. The current tape-based archival system is expensive (the Academy spends millions annually on film preservation), requires regular migration, and occupies significant physical space. DNA could store the entire history of cinema in a container the size of a shoe box.
Healthcare data archival is another target application. Modern genomic medicine generates enormous per-patient datasets (a whole genome sequence is approximately 200 GB), and regulatory requirements in many jurisdictions mandate retention for the patient’s lifetime plus 5-10 years. As genomic testing becomes routine, the cumulative archival data burden grows exponentially. DNA storage’s density and longevity align well with the healthcare archival use case.
The Timeline to Commercialization
Based on current cost trajectories and technology development, the DNA storage roadmap looks approximately like this: 2026-2028 will see continued cost reduction in DNA synthesis and the first limited commercial demonstrations of DNA archival storage for high-value, small-volume use cases (precious cultural archives, government intelligence archives). By 2028-2030, synthesis costs should reach the level where DNA competes with tape for the most expensive archival tiers (cold storage that’s retained for decades). By 2032-2035, DNA storage could become the default medium for long-term archival, with automated write-read systems integrated into data center infrastructure.
This timeline could accelerate if enzymatic synthesis achieves breakthrough cost reductions or if a major technology company (Google, Microsoft, Amazon) makes a large-scale production investment. It could decelerate if synthesis cost reductions plateau or if practical engineering challenges (automated handling of DNA molecules, quality control at scale, random access performance) prove harder to solve than anticipated. The science is proven; the engineering and economics are the remaining challenges — a pattern that historically resolves through sustained investment and iteration.
Related articles: Fintech Super Apps Dominate Emerging Mar | Neuromorphic Computing: Brain-Inspired C | 3D Bioprinting in 2026: From Lab Curiosi